Separable convolution: Part 2
Back in October I introduced the concept of filter separability. A two-dimensional filter s is said to be separable if it can be written as the convolution of two one-dimensional filters v and h:

I said then that "next time" I would explain how to determine whether a given filter is separable. Well, I guess I got side-tracked, but I'm back on topic now.
This question gave me one of earliest opportunities at The MathWorks to wander down to company co-founder Cleve's office and ask for advice. I asked, "How can I determine if a matrix is an outer product of two vectors?" Cleve was very helpful, as he always is, although I was a little embarrassed afterward that I hadn't figured it out myself. "Go look at the rank function," Cleve told me.
Of course. If a matrix is an outer product of two vectors, its rank is 1. Here are the key lines of code in rank:
dbtype 15:19 rank
15 s = svd(A); 16 if nargin==1 17 tol = max(size(A)') * eps(max(s)); 18 end 19 r = sum(s > tol);
So the test is this: The rank of A is the number of nonzero singular values of A, with some numerical tolerance based on eps and the size of A.
Let's try it with a few common filters.
An averaging filter should be obvious:
averaging = ones(5,5) / 25; rank(averaging)
ans =
1
The Sobel kernel:
sobel = [-1 0 1; -2 0 2; -1 0 1]; rank(sobel)
ans =
1
The two-dimensional Gaussian is the only radially symmetric function that is also separable:
gaussian = fspecial('gaussian');
rank(gaussian)
ans =
1
A disk is not separable:
disk = fspecial('disk');
rank(disk)
ans =
5
So how can we determine the outer product vectors? The answer is to go back to the svd function. Here's a snippet from the doc: [U,S,V] = svd(X) produces a diagonal matrix S of the same dimension as X, with nonnegative diagonal elements in decreasing order, and unitary matrices U and V so that X = U*S*V'.
A rank 1 matrix has only one nonzero singular value, so U*S*V' becomes U(:,1) * S(1,1) * V(:,1)'. This is basically the outer product we were seeking. Therefore, we want the first columns of U and V. (We have to remember also to use the nonzero singular value as a scale factor.)
Let's try this with the Gaussian filter.
[U,S,V] = svd(gaussian)
U =
-0.1329 0.9581 -0.2537
-0.9822 -0.1617 -0.0959
-0.1329 0.2364 0.9625
S =
0.6420 0 0
0 0.0000 0
0 0 0.0000
V =
-0.1329 -0.6945 -0.7071
-0.9822 0.1880 0.0000
-0.1329 -0.6945 0.7071
Now get the horizontal and vertical vectors from the first columns of U and V.
v = U(:,1) * sqrt(S(1,1))
v = -0.1065 -0.7870 -0.1065
h = V(:,1)' * sqrt(S(1,1))
h = -0.1065 -0.7870 -0.1065
I have chosen, somewhat arbitrarily to split the scale factor, S(1,1), "equally" between v and h.
Now check to make sure this works:
gaussian - v*h
ans = 1.0e-015 * -0.0243 -0.1527 -0.0243 -0.0139 -0.1110 -0.0139 -0.0035 0 -0.0035
Except for normal floating-point roundoff differences, gaussian and v*h are equal.
You can find code similar to this in the MATLAB function filter2, as well as in the Image Processing Toolbox function imfilter.




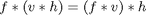




Comments
To leave a comment, please click here to sign in to your MathWorks Account or create a new one.