Deep learning deciphers what rats are saying
For many years, researchers knew that rodents’ squeaks tell a lot about how the animals are feeling. Much like a wagging tail on a dog, certain vocalizations indicate the rodents are happy. Conversely, other vocalizations indicate the rodents are stressed, or even depressed.
But why were they interested in the rodents’ moods? These researchers wanted to understand the rodents’ responses to various stimuli. This can help researchers determine the best way to help people who are addicted or depressed. They would be able to tell if a treatment helped reduce the feelings of depression by simply analyzing how the rodents chatted.
Rat chatter is difficult to decode since rodents communicate largely in ultrasonic vocalizations (USVs) that human ears cannot hear. USVs range from 20 kHz to 115 kHz, while humans can typically hear sounds from 20 Hz to 20 kHz.
Up until now, researchers have relied heavily on time-consuming, manual analysis of rodent chatter. The vocalizations are at such a high frequency, researchers had to slow down the recordings in order to hear them. Even with specialized microphones, tagging and categorizing the high-pitched squeaks in recordings is labor intensive. These methods are also vulnerable to human error and misinterpretation.
“In the past, researchers have recorded these to gain better insights into the emotional state of an animal during behavior testing,” Dr. John Neumaier, a professor in the Department of Psychiatry and Behavioral Sciences, told Digital Trends. “The problem was that manual analysis of these recordings could take 10 times longer to listen to when slowed down to frequencies that humans can hear. This made the workload exhaustive and discouraged researchers from using this natural read out about animals’ emotional states.”
So, the team of researchers from the University of Washington turned to artificial intelligence (AI) to automate the process. Their program is called DeepSqueak, since it relies on a form of AI called deep learning.
Using deep learning to analyze the USVs
Two researchers, Russell Marx, a technician in the Psychiatry & Behavioral Sciences at the University of Washington, and Dr. Kevin Coffey, a postdoctoral fellow at the University of Washington, worked with Professor Neumaier to create DeepSqueak, software that detects and analyzes USVs. Their research was recently published in the Nature Journal of Neuropsychopharmacology.
“We can train the software to analyze these calls in a way that is much more similar to how humans learn,” said Coffey. “Rather than mathematically describing what a vocalization is, we just show it pictures and examples.”
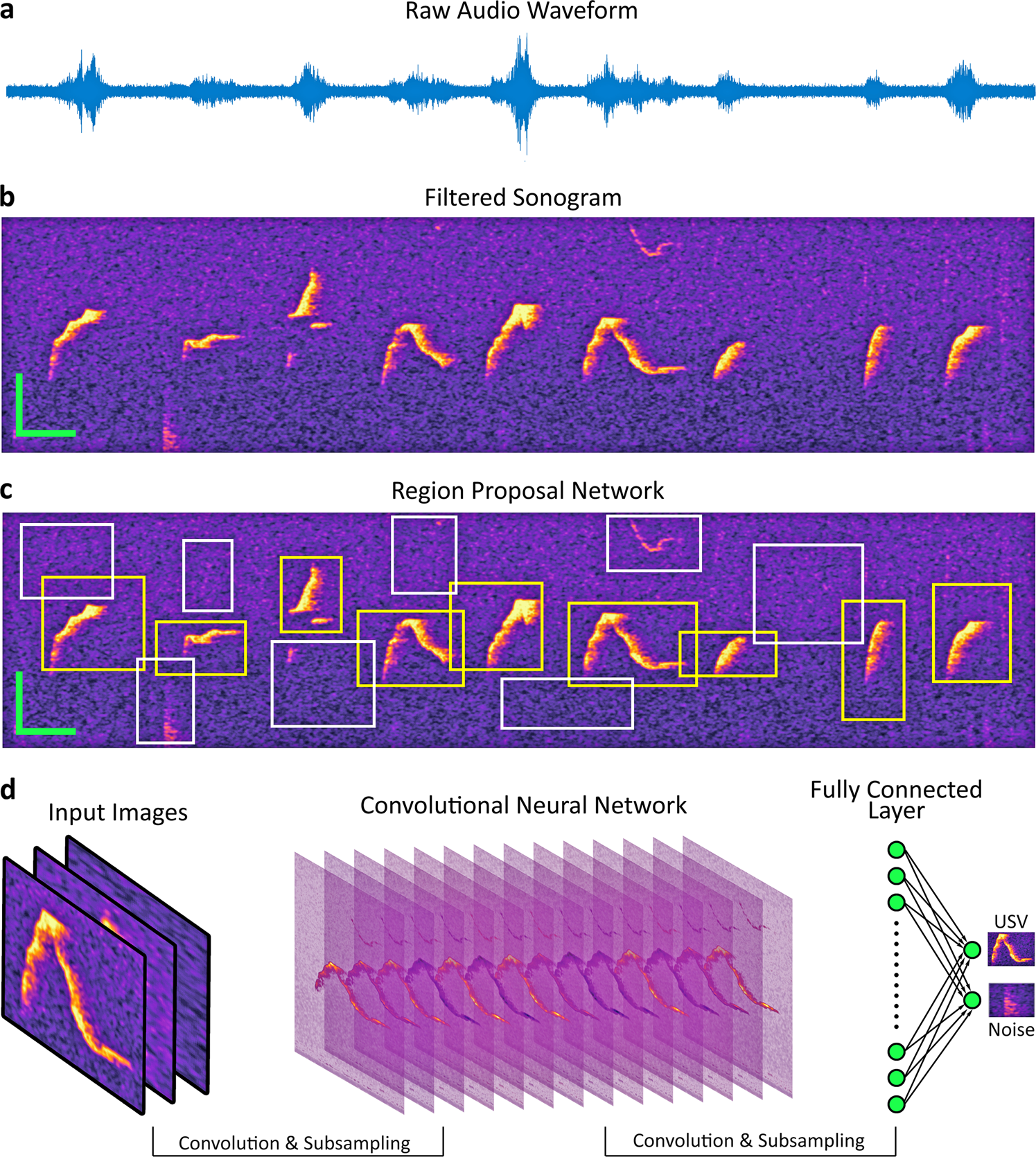
DeepSqueak works by turning an audio problem into a visual problem. The input to DeepSqueak is an audio file (.WAV or .FLAC). DeepSqueak splits the audio files into short segments and then converts these segments into images (sonograms). The figure below shows the transformation from a raw audio file to a filtered sonogram.
The sonograms are fed into a deep learning AI program that identifies and classifies the images, much like the AI used in self-driving cars to identify stop signs and lane markers. It first decides if a squeak is present in the sonogram, and if so, what type of squeak it is.
“DeepSqueak uses biomimetic algorithms that learn to isolate vocalizations by being given labeled examples of vocalizations and noise,” said Marx.
The team started DeepSqueak using example code, Object Detection Using Faster R-CNN Deep Learning from the MathWorks website. From there, they developed the DeepSqueak software package and GUI in MATLAB. DeepSqueak uses Computer Vision System Toolbox, Curve Fitting Toolbox, Image Processing Toolbox, Parallel Computing Toolbox, and Deep Learning Toolbox.
Technology can help develop better treatments for addiction
The research team is focused on psychiatry and behavioral science. This non-invasive research found that the rodents are happiest when anticipating a reward, such as sugar, or playing with their peers. They also found male rodents behaved differently when female rodents were around. No big surprise there.
Professor Neumaier says his goal is to develop treatments for stress disorders and addiction. DeepSqueak will help the lab get there much faster by making deciphering ultrasonic vocalizations convenient and quick.
“If scientists can understand better how drugs change brain activity to cause pleasure or unpleasant feelings, we could devise better treatments for addiction,” he said.
The team has made DeepSqueak available to all researchers so they can create their own analysis. The code is on Github. The program can currently identify approximately 20 different USVs. The team hopes that as others identify and tag various USVs, they’ll be able to create a virtual Google Translate for rat chatter.
To learn more about DeepSqueak, check out this video:








Comments
To leave a comment, please click here to sign in to your MathWorks Account or create a new one.