Matrix Condition and the QR Decomposition
The QR decomposition provides an estimate of the matrix condition number.
Contents
Query
Professor Magdy Hanna of the Department of Engineering Mathematics and Physics at Fayoum University in Fayoum, Egypt, asked me:
If we have the QR decomposition of a matrix, with or without pivoting, is there an easy way to get the condition number of the matrix without starting from scratch and using the SVD?
Short answer
The short answer to Prof. Hanna's question is:
Yes. The ratio of the largest and smallest diagonal elements of qr(A) is often a good estimate of cond(A).
Compare this estimate to the exact condition of a magic square. Produce a score measuring how well the estimate does.
A = magic(11);
R = qr(A);
d = abs(diag(R));
est = max(d)/min(d);
kappa = cond(A);
disp(" est kappa score")
disp([est kappa est/kappa])
est kappa score
2.6964 11.1021 0.2429
QR Decomposition
A more complete answer involves column pivoting.
The QR decomposition without column pivoting is an orthogonal Q and an upper triangular R so that
A = Q*R
The QR composition with column pivoting is an orthogonal Q, an upper triangular R and a permutation P that reorders the columns of A so that
A*P = Q*R
In either case, Q and P are orthogonal, so
cond(R) = cond(A)
Any estimate of the condition of R provides an estimate of the condition of A.
tricond
An estimate the condition of a triangular matrix is the ratio of the largest and smallest elements on the diagonal.
type tricond
function cond = tricond(T)
% Estimate condition of lower or upper triangular matrix.
if istril(T) || istriu(T)
D = diag(abs(T));
cond = max(D)/min(D);
else
cond = NaN;
end
end
qr function
Here is the 4-by-4 Pascal matrix
A = pascal(4)
A =
1 1 1 1
1 2 3 4
1 3 6 10
1 4 10 20
This matrix is reasonably well conditioned.
kappa = cond(A)
kappa = 691.9374
With one output, the qr function provides the "Q-less" QR decomposition with no pivoting.
(With two outputs, the qr function produces the same R and also reveals the orthogonal Q.)
R = qr(A)
R =
-2.0000 -5.0000 -10.0000 -17.5000
0 -2.2361 -6.7082 -14.0872
0 0 1.0000 3.5000
0 0 0 -0.2236
For this matrix tricond without pivoting produces a poor estimate of the exact condition.
trico = tricond(R);
kappa = cond(A);
disp(" tricond kappa score")
disp([trico kappa trico/kappa])
tricond kappa score 10.0000 691.9374 0.0145
With three outputs, qr does column pivoting and returns a different R.
[Q,R,P] = qr(A);
R
R =
-22.7376 -5.2336 -1.5393 -12.0065
0 -1.6153 -1.2034 -1.3387
0 0 0.4270 -0.2170
0 0 0 -0.0638
Compare tricond and the exact condition.
trico = tricond(R);
disp(" tricond kappa score")
disp([trico kappa trico/kappa])
tricond kappa score 356.6259 691.9374 0.5154
Column pivoting produces a much better tricond estimate for this matrix.
Gallery
Let's do more tests using these matrices from the MATLAB Gallery.
funs = ["moler", "parter", "binomial", "chebspec", "cauchy",... "chebspec", "chebvand","circul", "frank", "ris"];
One of these Gallery matrices is the "moler" matrix, even though I didn't invent it. A precursor to the "moler" matrix is a badly conditioned upper triangular matrix with ones on the diagonal, minus ones above the diagonal and zeros below.
n = 5;
U = eye(n) - triu(ones(n),1)
U =
1 -1 -1 -1 -1
0 1 -1 -1 -1
0 0 1 -1 -1
0 0 0 1 -1
0 0 0 0 1
The "moler" matrix is
M = U'*U
M =
1 -1 -1 -1 -1
-1 2 0 0 0
-1 0 3 1 1
-1 0 1 4 2
-1 0 1 2 5
The "moler" matrix squares the condition of U. The condition of M grows like 4^n.
kappa = cond(M)
kappa = 865.9801
The tricond score without column pivoting is very poor.
R = qr(M);
trico = tricond(R);
disp(" tricond kappa score")
disp([trico kappa trico/kappa])
tricond kappa score 20.7364 865.9801 0.0239
With pivoting the score is much better.
[~,R,~] = qr(M);
trico = tricond(R);
disp(" tricond kappa score")
disp([trico kappa trico/kappa])
tricond kappa score 555.0198 865.9801 0.6409
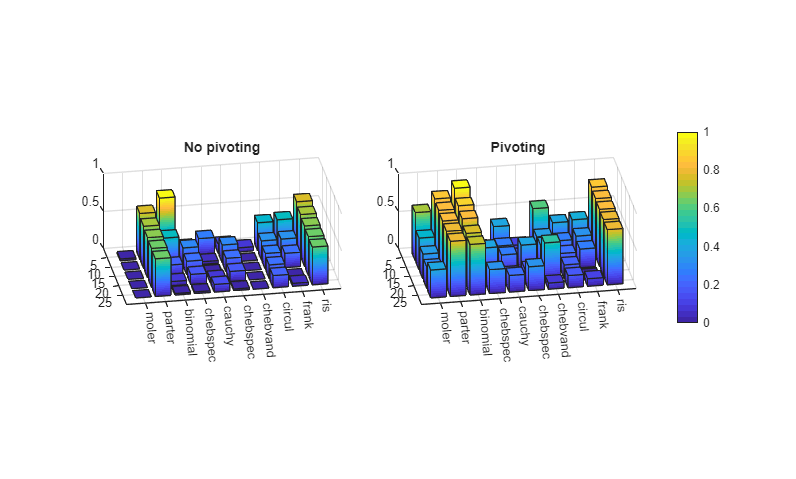
These scores are in the first line of the following output and in the upper left-hand corners of the 3-D bar graphs.
Tests
[Dnopiv,Dpiv,xt,yt] = tester(funs);
fun n nopiv piv
moler 5 0.024 0.641
10 0.000 0.426
15 0.000 0.337
20 0.000 0.287
25 0.000 0.372
parter 5 0.610 0.804
10 0.564 0.776
15 0.536 0.757
20 0.517 0.744
25 0.502 0.735
binomial 5 0.795 0.926
10 0.387 0.817
15 0.150 0.768
20 0.056 0.696
25 0.021 0.674
chebspec 5 0.120 0.110
10 0.236 0.095
15 0.183 0.358
20 0.132 0.165
25 0.022 0.318
cauchy 5 0.217 0.376
10 0.034 0.242
15 0.016 0.247
20 0.058 0.116
25 0.102 0.222
chebspec 5 0.120 0.110
10 0.236 0.095
15 0.183 0.358
20 0.132 0.165
25 0.022 0.318
chebvand 5 0.055 0.577
10 0.002 0.326
15 0.000 0.338
20 0.000 0.492
25 0.000 0.082
circul 5 0.370 0.370
10 0.252 0.252
15 0.209 0.209
20 0.180 0.180
25 0.162 0.162
frank 5 0.390 0.390
10 0.257 0.296
15 0.191 0.248
20 Inf 0.013
25 0.039 0.100
ris 5 0.610 0.804
10 0.564 0.776
15 0.536 0.757
20 0.517 0.744
25 0.502 0.735
Plot
[c,ax1,ax2] = ploter(funs,Dnopiv,Dpiv,xt,yt);







评论
要发表评论,请点击 此处 登录到您的 MathWorks 帐户或创建一个新帐户。