For Bioinformaticists…or GUI Builders
Note
The file submission referenced in this post is no longer available on File Exchange.
This week's Pick: Primer Design Tool, by Brian Madsen.
If you work in the field of bioinformatics, and especially if you do any primer design, you might be aware of a really nice demo that ships with the Bioinformatics Toolbox, and that shows how to use our tools for that task.
The script for primerdemo steps through the importing of data (from GenBank, for instance); calculating oligonucleotide properties; finding potential forward and reverse primers, and filtering them based on (for example) GC content, melting temperature, and dimerization; and visualizing and selecting primers based on user-specified criteria. This is all great stuff, if you are a bioinformaticist.
Brian's "Primer Design Tool" takes all of these steps and wraps them very neatly into a GUI environment that makes it very easy to modify constraints and to see the results of changing criteria. This is a very nice tool for folks who do primer design.
However, what I really like about Brian's file is that it is a very useful demonstration of how converting code-based functionality to GUI-based functionality can, with a little bit of effort, allow you to leverage the great demos that ship with our products. I don't personally do a lot of primer design, for instance, but henceforth, whenever I am called upon to show this capability, I show the in-product demo, and then I show Brian's GUI implementation of the functionality.
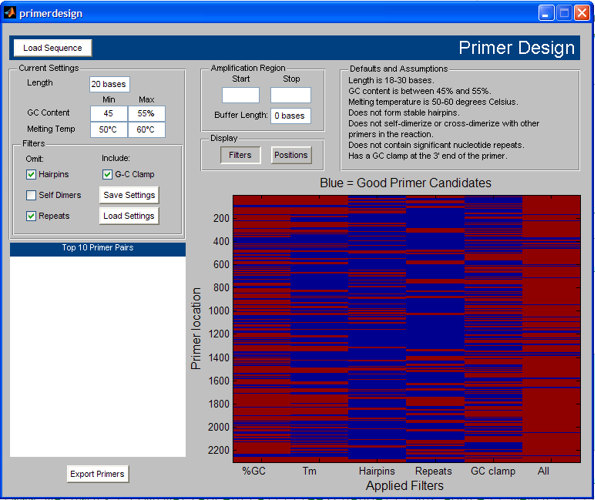
This is good stuff, even if you have no idea what an oligonucleotide is, or what primer design is all about.
Comments? Share them here.
- Category:
- Picks









Comments
To leave a comment, please click here to sign in to your MathWorks Account or create a new one.