Analyzing Fitness Data from Wearable Devices in MATLAB
Collecting and tracking health and fitness data with wearable devices is about to go mainstream as the smartphone giants like Apple, Google and Samsung jump into the fray. But if you collect data, what's the point if you don't analyze it?
Today's guest blogger, Toshi Takeuchi, would like to share an analysis of a weight lifting dataset he found in a public repository.
Contents
- Motivation, dataset, and prediction accuracy
- Data preprocessing and exploratory analysis
- Predictive Modeling with Random Forest
- Plot misclassification errors by number of trees
- Variable Importance
- Evaluate trade-off with ROC plot
- The reduced model with 12 features
- Conclusion and the next steps - integrate your code into your app
Motivation, dataset, and prediction accuracy
The Human Activity Recognition (HAR) Weight Lifting Exercise Dataset provides measurements to determine "how well an activity was performed". 6 subjects performed 1 set of 10 Unilateral Dumbbell Biceps Curl in 5 different ways.
When I came across this dataset, I immediately thought of building a mobile app to advise end users whether they are performing the exercise correctly, and if not, which common mistakes they are making. I used the powerful 'Random Forest' algorithm to see if I could build a successful predictive model to enable such an app. I was able to achieve 99% prediction accuracy with this dataset and I would like to share my results with you.
The dataset provides 39,242 samples with 159 variables labeled with 5 types of activity to detect - 1 correct method and 4 common mistakes:
- exactly according to the specification (Class A)
- throwing the elbows to the front (Class B)
- lifting the dumbbell only halfway (Class C)
- lowering the dumbbell only halfway (Class D)
- throwing the hips to the front (Class E)
Sensors were placed on the subjects' belts, armbands, glove and dumbbells, as described below:
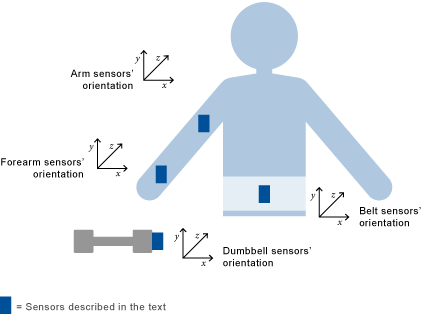
Citation Velloso, E.; Bulling, A.; Gellersen, H.; Ugulino, W.; Fuks, H. Qualitative Activity Recognition of Weight Lifting Exercises. Proceedings of 4th International Conference in Cooperation with SIGCHI (Augmented Human '13) . Stuttgart, Germany: ACM SIGCHI, 2013. Read more:
Data preprocessing and exploratory analysis
Usually you cannot use raw data directly. Preprocessing is an important part of your analysis workflow that has significant downstream impact.
- Load the dataset and inspect data for missing values
- Partition the dataset for cross validation
- Clean and normalize variables
- Select predictor variables (features)
Among those steps, cross validation is a key step specific to predictive modeling. Roughly speaking, you hold out part of available data for testing later, and build models using the remaining dataset. The held out set is called the 'test set' and the set we use for modeling is called the 'training set'. This makes it more difficult to overfit your model, because you can test your model against the data you didn't use in the modeling process, giving you a realistic idea how the model would actually perform with unknown data.
Exploratory analysis usually begins with visualizing data to get oriented with its nuances. Plots of the first four variables below show that:
- data is organized by class - like 'AAAAABBBBBCCCCC'. This can be an artifact of the way the data was collected and real life data may not be structured like this, so we want to use more realistic data to build our model. We can fix this issue by randomly reshuffling the data.
- data points cluster around a few different mean values - indicating that measurements were taken by devices calibrated in a few different ways
- those variables exhibit a distinct pattern for Class E (colored in magenta) - those variables will be useful to isolate it
Review preprocess.m if you are curious about the details.
preprocess

Predictive Modeling with Random Forest
The dataset has some issues with calibration. We could further preprocess the data in order to remove calibration gaps. This time, however, I would like to use a flexible predictive algorithm called Random Forest. In MATLAB, this algorithm is implemented in the TreeBagger class available in Statistics Toolbox.
Random Forest became popular particularly after it was used by a number of winners in Kaggle competitions. It uses a large ensemble of decision trees (thus 'forest') trained on random subsets of data and uses the majority votes of those trees to predict the result. It tends to produce a highly accurate result, but the complexity of the algorithm makes it slow and difficult to interpret.
To accelerate the computation, I will enable the parallel option supported by Parallel Computing Toolbox. You can comment out unnecessary code if you don't use it.
Once the model is built, you will see the confusion matrix that compares the actual class labels to predicted class labels. If everything lines up on a diagonal line, then you achieved 100% accuracy. Off-diagonal numbers are misclassification errors.
The model has a very high prediction accuracy even though we saw earlier that our dataset had some calibration issues.
Initialize parallel option - comment out if you don't use parallel
poolobj = gcp('nocreate'); % don't create a new pool even if no pool exits if isempty(poolobj) parpool('local',2); end opts = statset('UseParallel',true);
Starting parallel pool (parpool) using the 'local' profile ... connected to 2 workers.
Create a Random Forest model with 100 trees, parallel enabled...
rfmodel = TreeBagger(100,table2array(Xtrain),Ytrain,... 'Method','classification','oobvarimp','on','options',opts);
Here's the non-parallel version of the same model
rfmodel = TreeBagger(100,table2array(Xtrain),Ytrain,...
'Method','classification','oobvarimp','on');Predict the class labels for the test set.
[Ypred,Yscore]= predict(rfmodel,table2array(Xtest));
Compute the confusion matrix and prediction accuracy.
C = confusionmat(Ytest,categorical(Ypred)); disp(array2table(C,'VariableNames',rfmodel.ClassNames,'RowNames',rfmodel.ClassNames)) fprintf('Prediction accuracy on test set: %f\n\n', sum(C(logical(eye(5))))/sum(sum(C)))
A B C D E
____ ___ ___ ___ ___
A 1133 1 0 0 0
B 4 728 1 0 0
C 0 3 645 3 0
D 0 0 8 651 0
E 0 0 0 6 741
Prediction accuracy on test set: 0.993374
Plot misclassification errors by number of trees
I happened to pick 100 trees in the model, but you can check the misclassification errors relative to the number of trees used in the prediction. The plot shows that 100 is an overkill - we could use fewer trees and that will make it go faster.
figure plot(oobError(rfmodel)); xlabel('Number of Grown Trees'); ylabel('Out-of-Bag Classification Error');

Variable Importance
One major criticism of Random Forest is that it is a black box algorithm and it not easy to understand what it is doing. However, Random Forest can provide the variable importance measure, which corresponds to the change in prediction error with and without the presence of a given variable in the model.
For our hypothetical weight lifting trainer mobile app, Random Forest would be too cumbersome and slow to implement, so you want to use a simpler prediction model with fewer predictor variables. Random Forest can help you with selecting which predictors you can drop without sacrificing the prediction accuracy too much.
Let's see how you can do this with TreeBagger.
Get the names of variables
vars = Xtrain.Properties.VariableNames;
Get and sort the variable importance scores. Because we turned 'oobvarimp' to 'on', the model contains OOBPermutedVarDeltaError which acts as the variable importance measure.
varimp = rfmodel.OOBPermutedVarDeltaError'; [~,idxvarimp]= sort(varimp); labels = vars(idxvarimp);
Plot the sorted scores.
figure barh(varimp(idxvarimp),1); ylim([1 52]); set(gca, 'YTickLabel',labels, 'YTick',1:numel(labels)) title('Variable Importance'); xlabel('score')

Evaluate trade-off with ROC plot
Now let's do the trade-off between the number of predictor variables and prediction accuracy. The Receiver operating characteristic (ROC) plot provides a convenient way to visualize and compare performance of binary classifiers. You plot the false positive rate against the true positive rate at various prediction thresholds to produce the curves. If you get a completely random result, the curve should follow a diagonal line. If you get a 100% accuracy, then the curve should hug the upper left corner. This means you can use the area under the curve (AUC) to evaluate how well each model performs.
Let's plot ROC curves with different sets of predictor variables, using the "C" class as the positive class, since we can only do this one class at a time, and the previous confusion matrix shows more misclassification errors for this class than others. You can use perfcurve to compute ROC curves.
Check out myROCplot.m to see the details.
nFeatures = [3,5,10,15,20,25,52];
myROCplot(Xtrain,Ytrain,Xtest,Ytest,'C',nFeatures,varimp)

The reduced model with 12 features
Based on the previous analysis, it looks like you can achieve a high accuracy rate even if you use as few as 10 features. Let's say we settled for 12 features. We now know you don't have to use the data from the glove for prediction, so that's one less sensor our hypothetical end users would have to buy. Given this result, I may even consider dropping the arm band, and just stick with the belt and dumbbell sensors.
Show which features were included.
disp(table(varimp(idxvarimp(1:12)),'RowNames',vars(idxvarimp(1:12)),... 'VariableNames',{'Importance'}));
Importance
__________
accel_belt_y 0.69746
pitch_dumbbell 0.77255
accel_arm_z 0.78941
accel_belt_x 0.81696
magnet_arm_y 0.81946
accel_arm_x 0.87168
magnet_arm_x 0.87897
accel_dumbbell_x 0.92222
magnet_forearm_x 1.0172
total_accel_belt 1.0461
gyros_arm_z 1.1077
gyros_belt_x 1.1235
Shut down the parallel pool.
delete(poolobj);
Conclusion and the next steps - integrate your code into your app
Despite my initial misgivings about the data, we were able to maintain high prediction accuracy with a Random Forest model with just 12 features. However, Random Forest is probably not an ideal model to implement on a mobile app given its memory foot print and slow response time.
The next step is to find a simpler model, such as logistics regression, that can perform decently. You may need to do more preprocessing of the data to make it work.
Finally, I have never tried this before, but you could generate C code out of MATLAB to incorporate it into a mobile app. Watch this webinar, MATLAB to iPhone Made Easy, for more details.

Do you track your activities with wearable devices? Have you tried analyzing the data? Tell us about your experience here!
- Category:
- Fun,
- Hardware,
- How To,
- Social Computing









Comments
To leave a comment, please click here to sign in to your MathWorks Account or create a new one.