Labeling Data Points
Jiro‘s pick this week is labelpoints by Adam Danz.
This week’s entry caught my attention for two reasons. One is that this entry does the task that I usually dread doing, which is making finishing touches to my plots. When I want to label some data points, I use the text function. It gives me some control of how to align the text, but it is basically limited to the extent of the text. For example, I can do this
x = [0 0 0];
y = [.8 .7 .6];
labels = {'label 1','label 2','label 3'};
plot(x,y,'o')
text(x,y,labels,'VerticalAlignment','bottom','HorizontalAlignment','right')
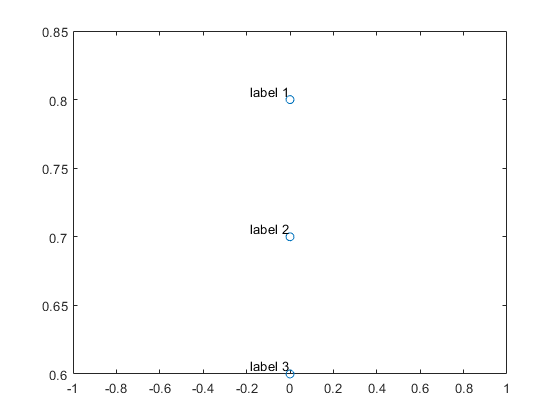
Or this.
plot(x,y,'o') text(x,y,labels,'VerticalAlignment','top','HorizontalAlignment','left')

But both of these aren’t exactly what I want because the labels slightly overlap the data. Also in the second one, the label goes beyond the bounds of the axes.
Usually at this point, I fiddle around with the coordinates of the text placements. Very doable, but tedious.
This is where Adam’s entry comes into play.
plot(x,y,'o') labelpoints(x,y,labels,'SE',0.2,1)

'SE' means southeast placement, 0.2 refers to the offset for the labels, and 1 means “adjust the x/y limits of the axes”. Perfect!
He also includes a wealth of options, one of which I found interesting being the option to detect outliers and only placing labels on the outliers. By making use of the Statistics and Machine Learning Toolbox, he provides different methods for detecting outliers,
x = 0:0.01:1; y = (0:0.01:1)+rand(1,101); labs = 1:101; plot(x,y,'o') labelpoints(x,y,labs,'outliers_lin',{'sd', 1.5})

The other reason Adam’s entry caught my attention was the amount of help and information he included in the entry. The first 300 lines of his code are help comments!! He includes many examples to test out all of the various options. As you can see from the File Exchange page, he has been keeping the file up-to-date periodically. I can tell he has put in a lot of time and effort into making and maintaining this code. I’m grateful that he has shared this with the community. Thanks Adam!
Comments
Give it a try and let us know what you think here or leave a comment for Adam.
- Category:
- Picks
 Cleve’s Corner: Cleve Moler on Mathematics and Computing
Cleve’s Corner: Cleve Moler on Mathematics and Computing The MATLAB Blog
The MATLAB Blog Guy on Simulink
Guy on Simulink MATLAB Community
MATLAB Community Artificial Intelligence
Artificial Intelligence Developer Zone
Developer Zone Stuart’s MATLAB Videos
Stuart’s MATLAB Videos Behind the Headlines
Behind the Headlines File Exchange Pick of the Week
File Exchange Pick of the Week Hans on IoT
Hans on IoT Student Lounge
Student Lounge MATLAB ユーザーコミュニティー
MATLAB ユーザーコミュニティー Startups, Accelerators, & Entrepreneurs
Startups, Accelerators, & Entrepreneurs Autonomous Systems
Autonomous Systems Quantitative Finance
Quantitative Finance MATLAB Graphics and App Building
MATLAB Graphics and App Building






Comments
To leave a comment, please click here to sign in to your MathWorks Account or create a new one.